CohesinDB v1.5 update: (start from Jan 1, 2026)
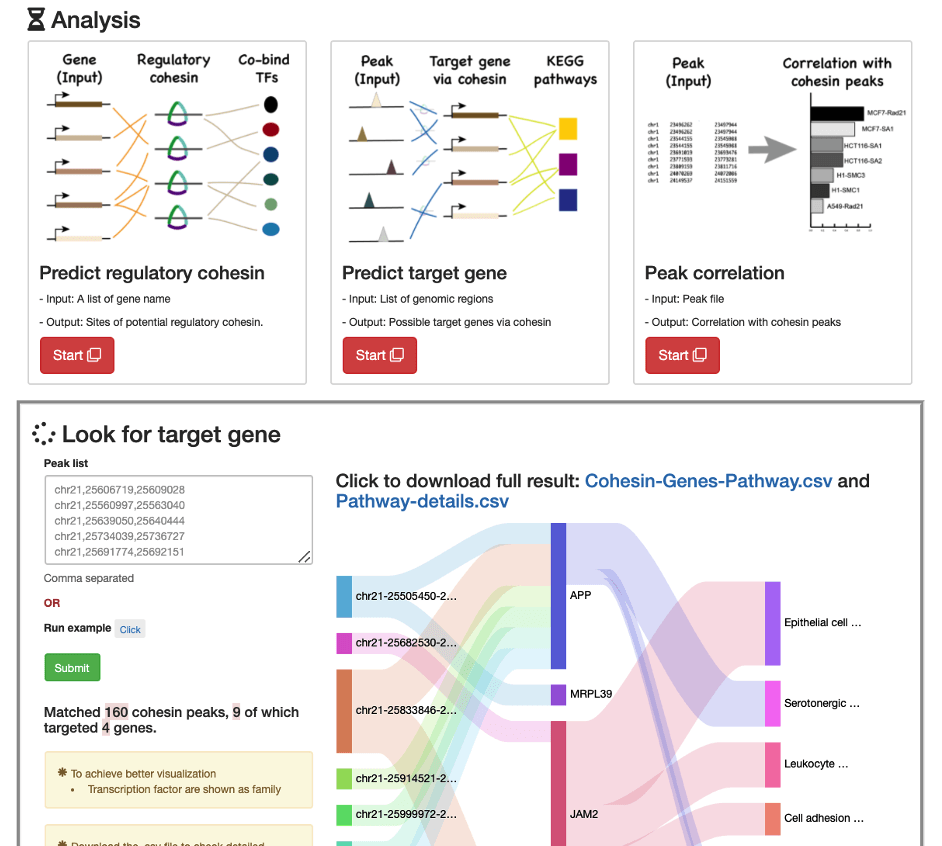
Building upon our previously developed multiomics resource CohesinDB, we mapped TF binding to cohesin-associated regulatory sites and linked these sites to their downstream target genes. The resulting Cohesin-GRN module in CohesinDB comprises 61,222,502 TF–cohesin site links and 2,228,634 cohesin site–gene links, collectively forming over 270 million TF–site–gene regulatory paths.
By enabling a CRE-informed and site-aware representation of gene regulation, Cohesin-GRN bridges conventional TF–gene GRNs with regulatory site-centric mechanisms. Given the pervasive roles of cohesin in enhancer activity, transcriptional control, and human disease, Cohesin-GRN provides a valuable resource for exploring transcriptional dysregulation and gene regulatory networks.
Please check the related publication about this update: Citation 2026
CohesinDB old version 2022~2025:
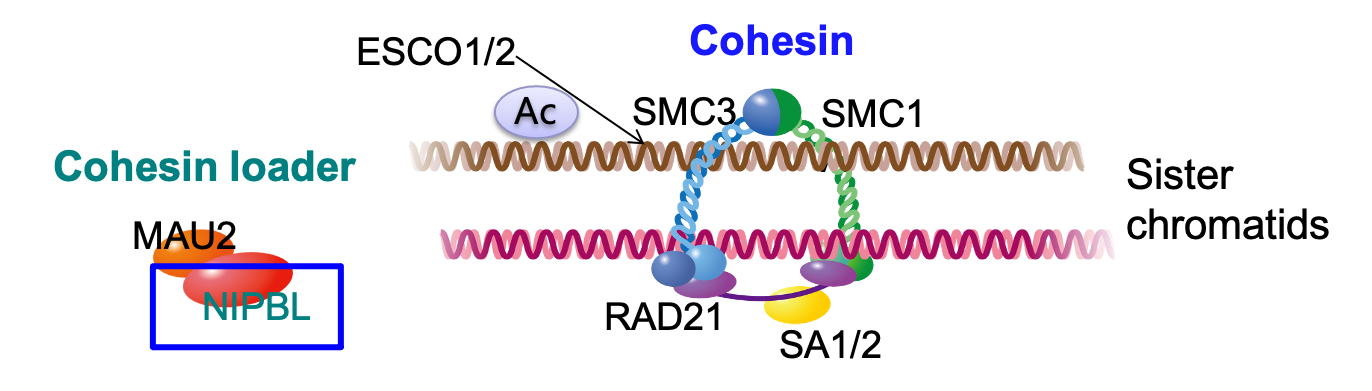
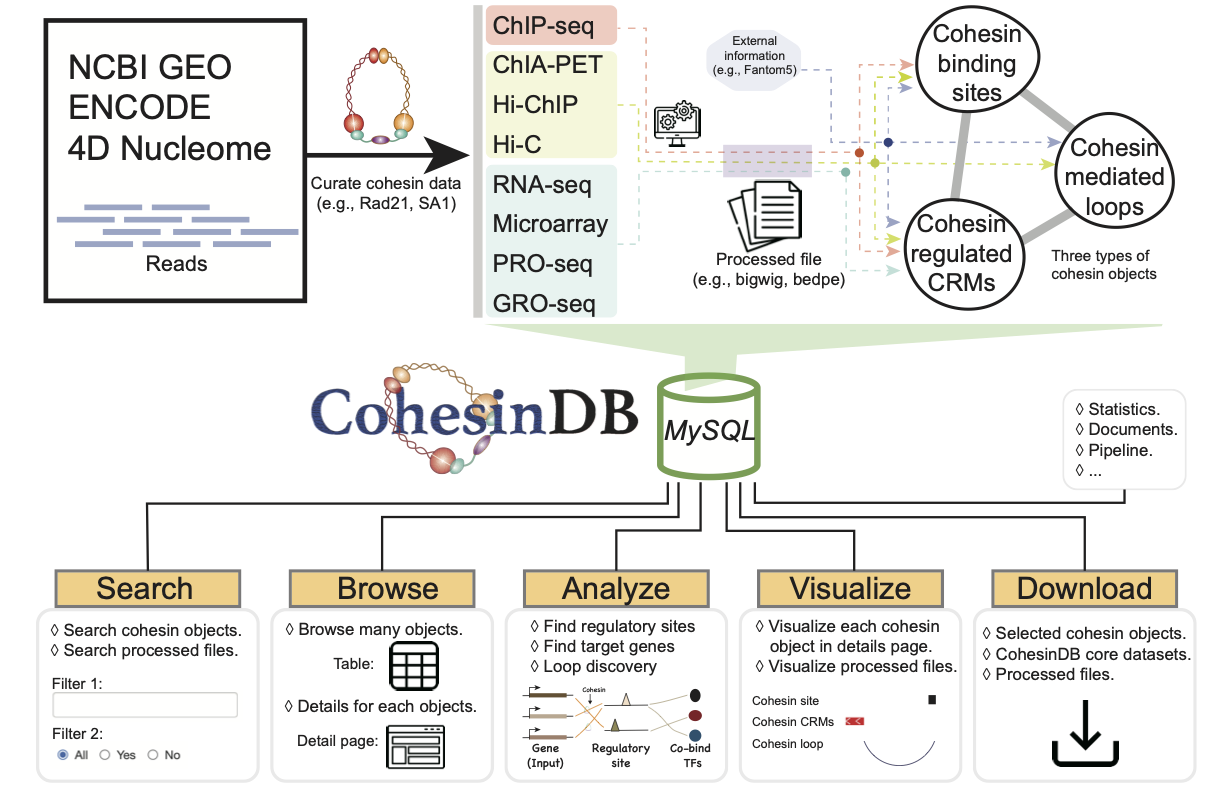
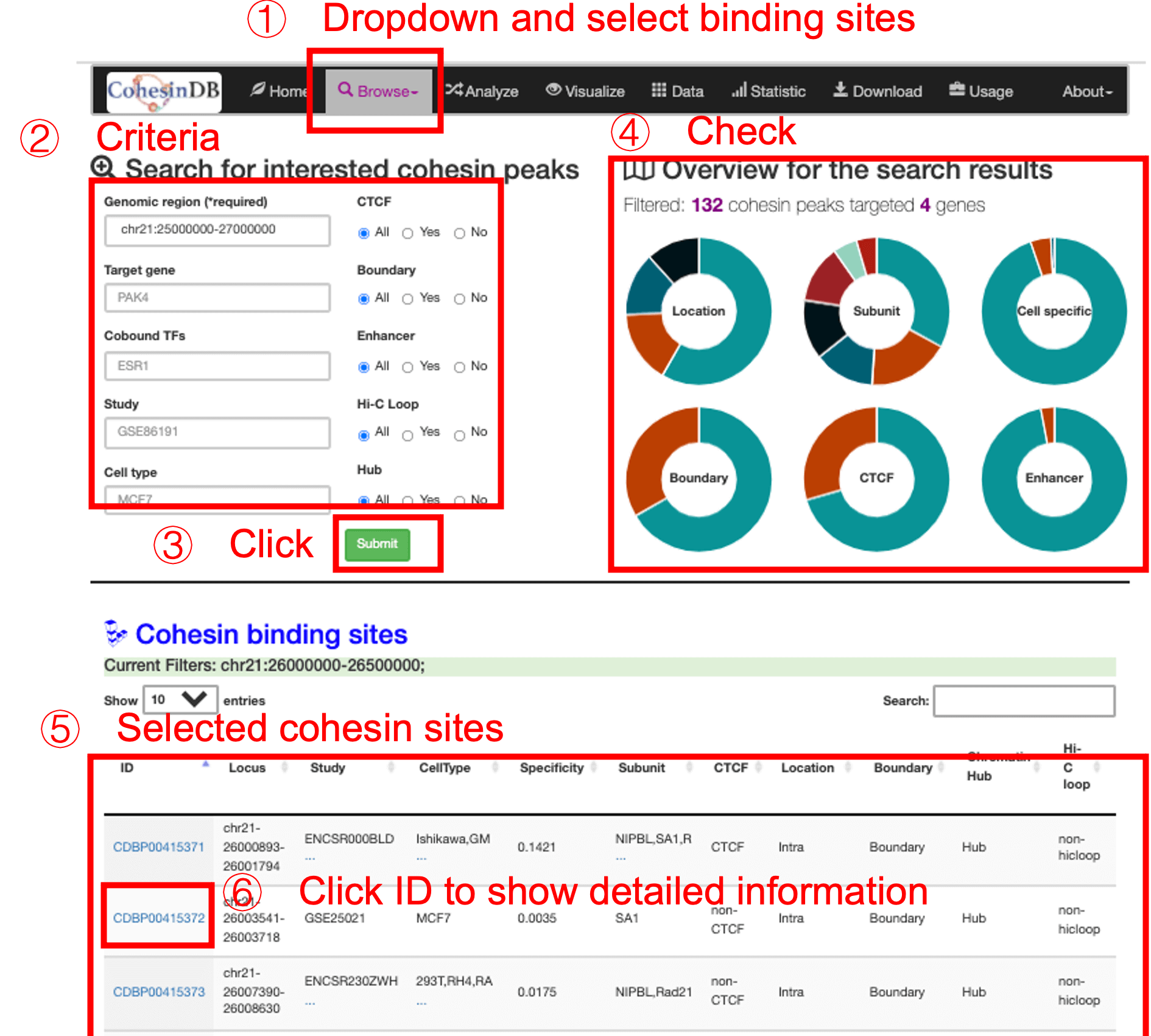
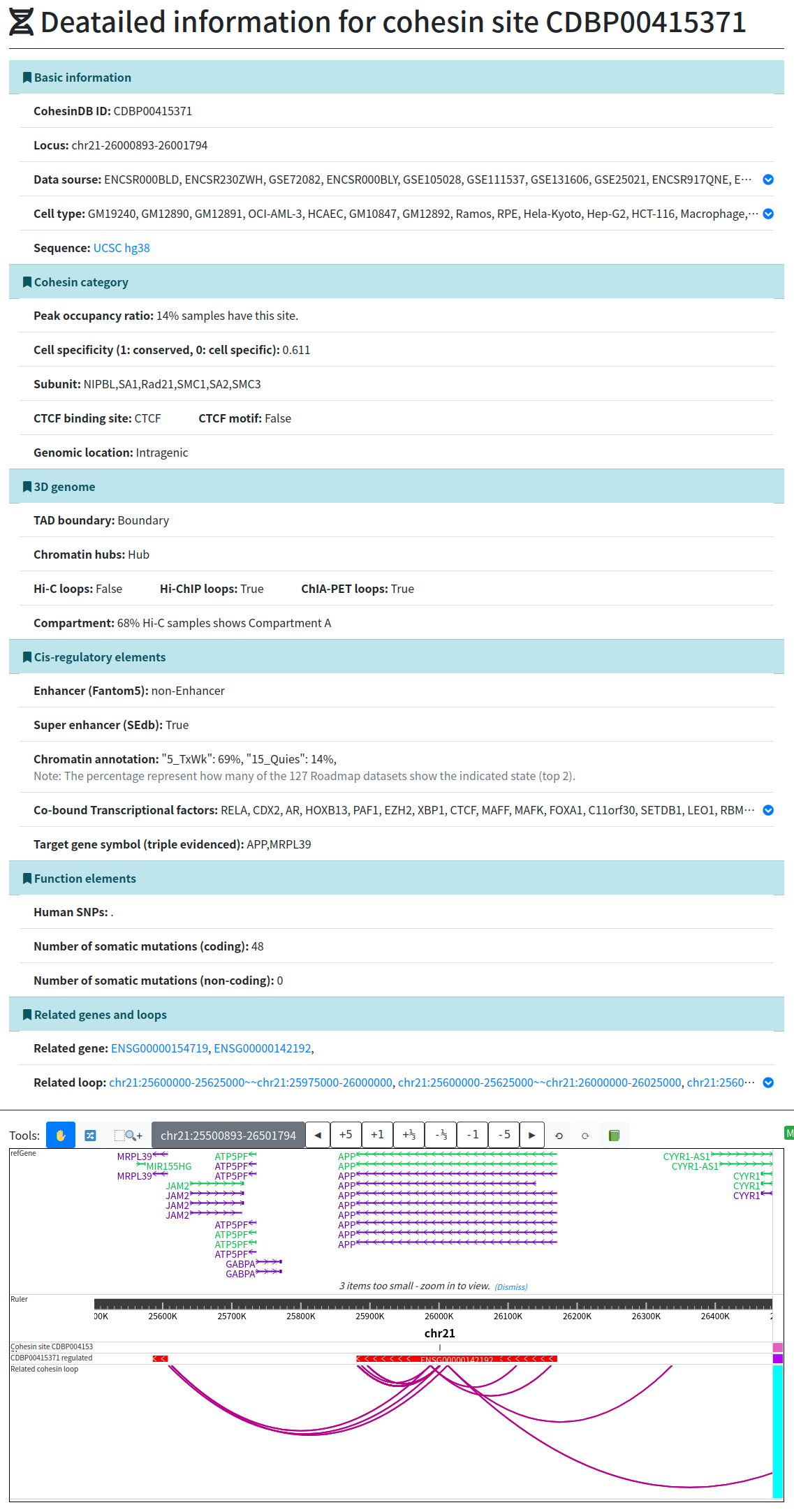
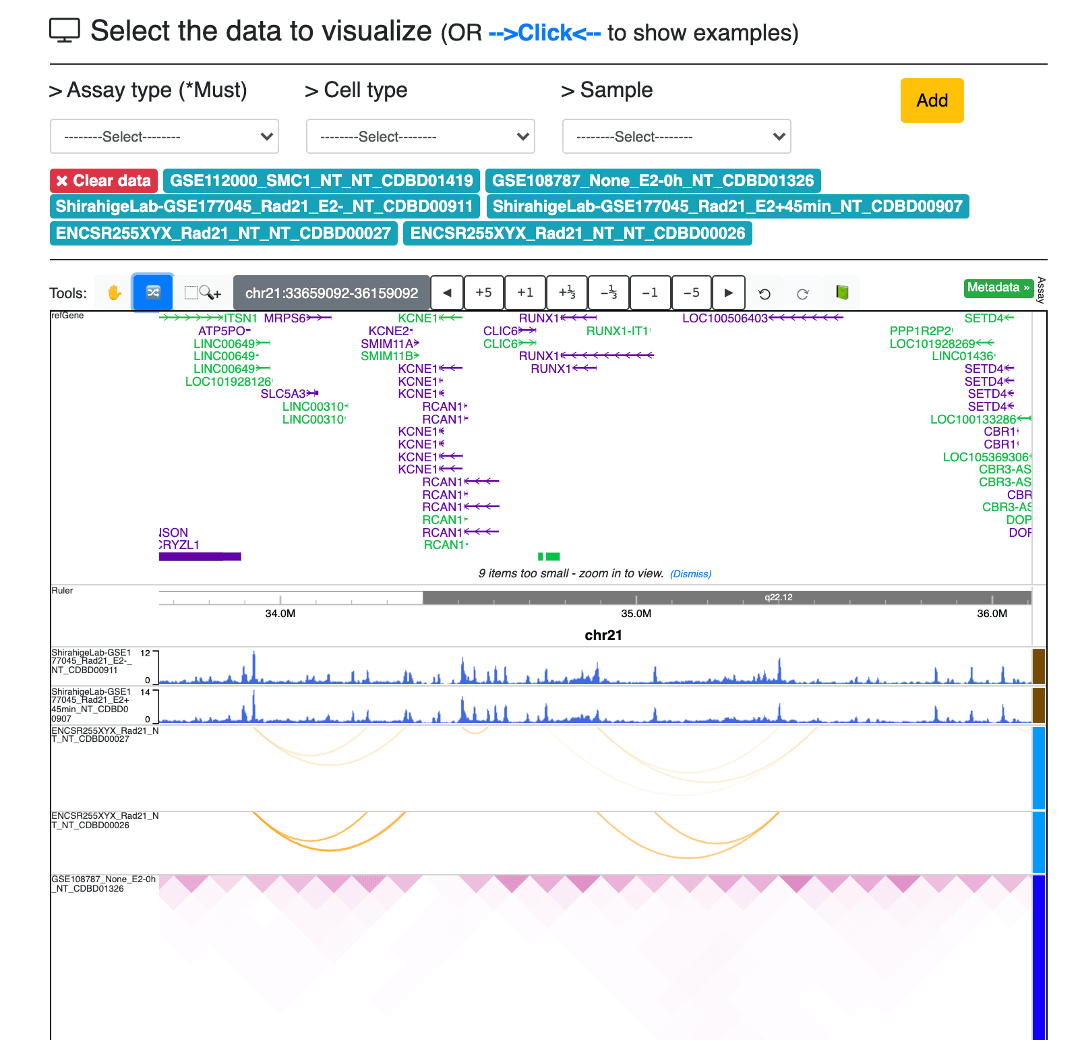
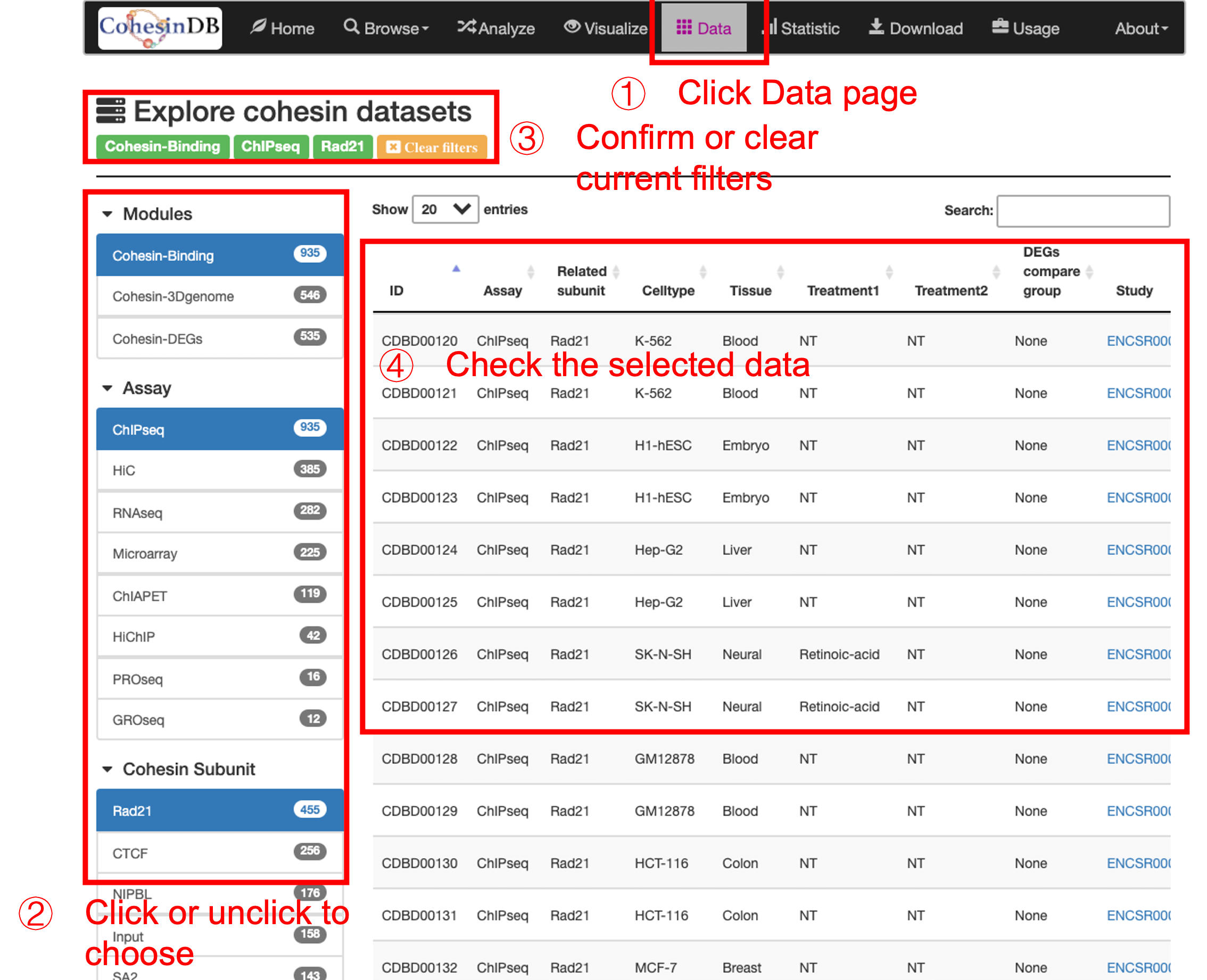
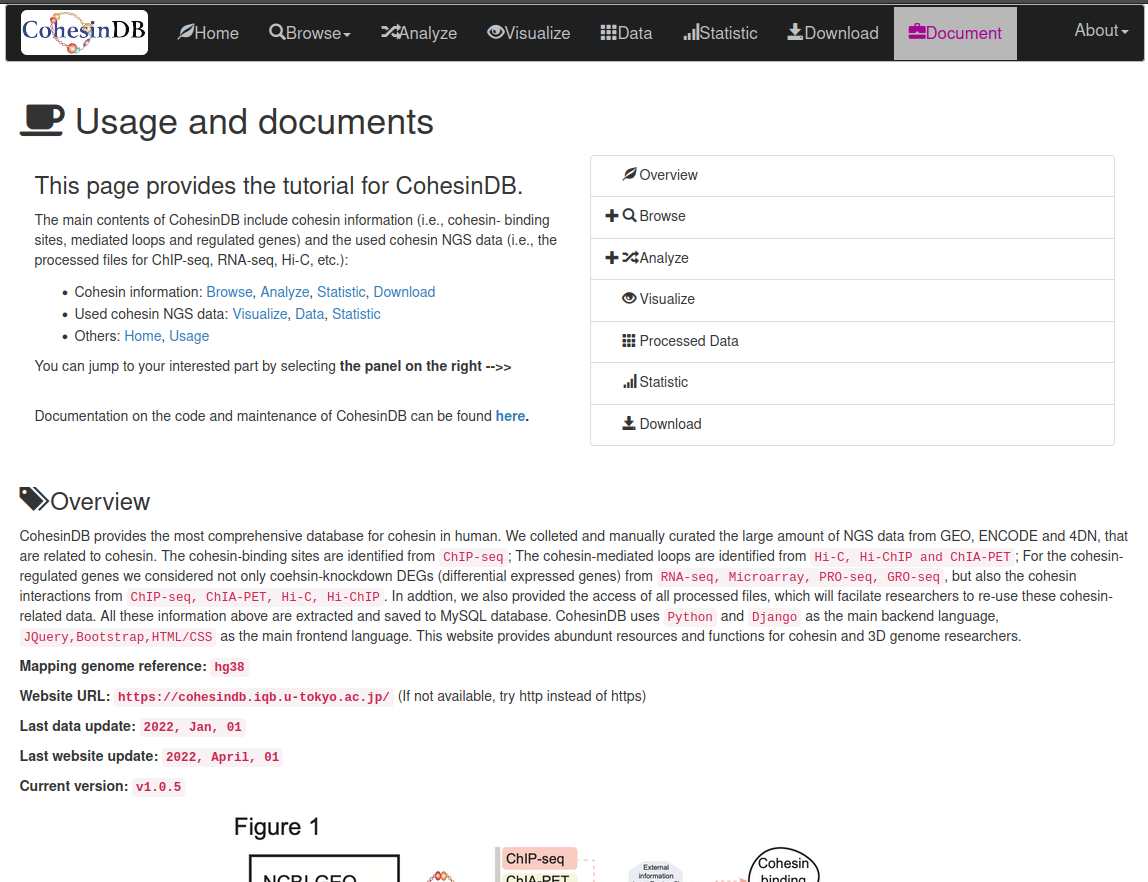
Welcome to CohesinDB (v1.2), a comprehensive database for decoding cohesin-related epigenomes, 3D genomes and transcriptomes in human cells. Cohesin is a multifunctional protein complex for transcriptional regulation and chromatin organization. By integrating multiomics NGS datasets, CohesinDB provides search and browse functions for: Cohesin-binding sites, Cohesin-mediated loops and Cohesin-regulated CRMs/genes. CohesinDB also have functions including Analyze, Visulize, Data, Statistic, Download and Document. CohesinDB is a valuable resource for cohesin, epigenomes, transcriptional regulation and chromatin organization.
If you use CohesinDB, please cite our NAR paper: Citation.